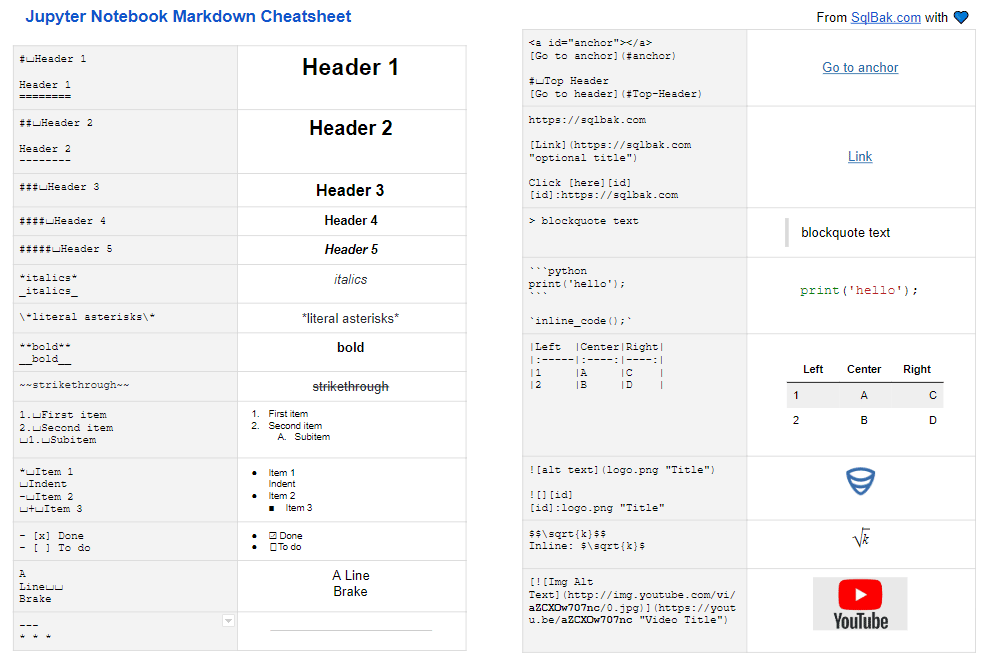
One can use the command summarize() to automatically produce summary tables for either numerical variables (i.e., variables where is.numeric() is TRUE) or categorical variables (where is.factor() is TRUE). # "fissure distance (mm)" "age (years)" "Subject" Now, the labels are labels(Orthodont) # distance age Subject We see that by setting variable labels, we also add the class 'ldf' to the data frame. If we now query if Orthodont is a labeled data frame and extract the labels, we get is.ldf(Orthodont) # TRUE class(Orthodont) # "ldf" "nfnGroupedData" "nfGroupedData" "groupedData" We use some of the information which is given on the help page of the Orthodont data and use it as labels: labels(Orthodont) <- c("fissure distance (mm)", "age (years)", "Subject", "Sex") 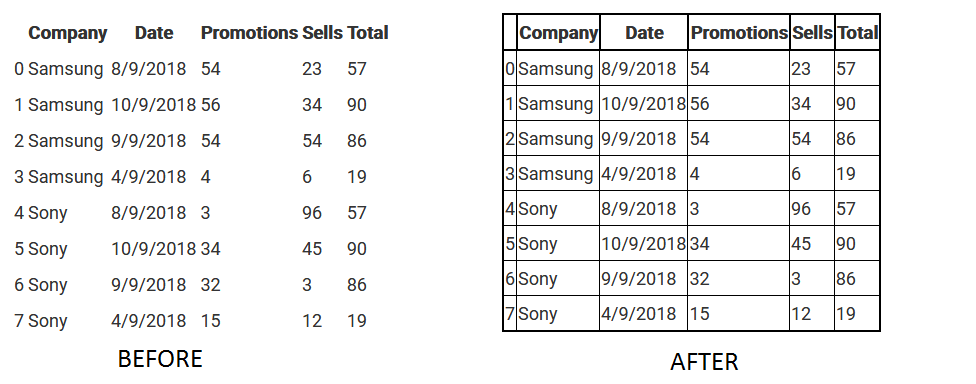
To explicitly set labels, which are usually more descriptive than the variable names, we can simply assign a vector of labels. This is a convenient feature, as we thus can relly on the fact that we will always have some variable labels. In this case, we simply get the variable names as no labels were set so far labels(Orthodont) # distance age Subject Sex To check if the data set is a labeled data set (i.e., of class 'ldf'), we can use is.ldf(Orthodont) # FALSEĭespite the fact that we do not have a labeled data frame, we can query the labels. # keep the original data set for later use First load the data data(Orthodont, package = "nlme") We use the Orthodont data package nlme throughout this tutorial. If we create a new ame we can extract and set variable labels using the function labels().
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
